Even though autism undoubtedly has biological underpinnings and the brains of those with autism may share some form of common structures (neuroanatomy) or function (neurophysiology), we still don't have a good working definition for it - or at least, nothing that everyone can agree upon. In the field of cancer research, even though scientists disagree about the primary causes of a tumour developing, they can all generally agree that uncontrolled cellular growth is a good working definition. If you see a tumour, you know it's following some sort of poorly-regulated growth process, even when it's benign (non-cancerous). Which makes finding causes of tumour promotion considerably easier. If you know the end result in terms of cell biology, that limits the number of variables you're searching for as instigators of that process.
But not so for autism. Even though the well-known psychiatrist, Leo Kanner, described the condition well over 70 years ago, the only thing people can seem to agree upon is a behavioural definition. (And even still, we argue over that!) We have done so much research and have published so many papers reporting correlations with things that may influence the occurrence of autism, ranging from the presence of one or more other disorders or diseases, down to associated genes. But unlike cancer, we have no general definition for what defines autism at the level of the cell nor at the level of the brain. We're still wandering around our research somewhat blindly.
Ultimately, that means we study just about everything that appears to have an association with the condition without much understanding of whether the relationship is causal or simply correlational in nature. An excellent example of this comes from studying autism genetics. Over 3,000 genes have found to have minor-to-strong association with the condition, both those which currently have no known cause (idiopathic) and conditions which are characterized by the association of several clinically recognizable features which tend to go together (syndromic), such as Fragile X Syndrome. While all roads may have once led to Rome, not all genes lead to autism. Those thousands of genes represent at least 12% of the genes within the human genome, a staggering proportion. It's highly unlikely that all or most converge onto a single behavioural trait.
just because two things occur frequently together that doesn't mean that one necessarily causes the other. As the hilarious statistical parody teaches us, the loss of pirates does not cause global warming even though the number of pirates have indeed declined over the last 130 years as global temperatures have simultaneously gone up.
You might wonder what pirates have to do with autism: what this analogy illustrates is that just because a gene mutation occurs more frequently in a particular sub-population doesn't necessarily mean that the gene has caused the condition of interest. This is especially true in the case of cancer. Some of the genes that mutate in cancer do play a role in its progression, but some gene mutations that are associated with the condition just occur because the disease itself makes it more likely for the genetic material in an organism to mutate - it causes genetic instability3.
This is called getting a "false positive", and in fact a recent article in Nature4 just dealt with this exact problem. The researchers reported that some genes which are less stable are more likely to mutate in cancer and therefore appear "associated" even though they are not causally related to the develop of the cancer itself. The problem is when geneticists assume that all genes mutate at the same rate, which is an incorrect assumption. Some genes are more likely to mutate than others, and rates can even vary by person. So the approach to studying mutations requires a more nuanced approach.
As mentioned, thousands of genes have been associated with autism, with approximately 100-200 genes considered "strongly associated". Interestingly, some of the same genes which the Nature paper determined were probably false positives in cancer studies and had nothing to do with the development of the condition are also high up on the autism list. These genes in particular are known for their high mutation rates. Coincidence?
The general point is that cancer researchers have realized that they may be receiving false positives in their genetics studies and are beginning to adjust their methods of analysis to account for this. Given the overlap in important genes associated with both cancer and autism, especially those genes which normally have high mutation rates, autism researchers may need to apply similar methods to whittle down results to those genes most likely play a role in the development of the condition.
Where do scientists go from here? Good question. I'm not absolutely certain, but one thing I do know is that in terms of research into the cause of autism, we need to make it our #1 priority to define autism at the cellular and tissue levels. Without this, we have no measuring stick for the feasibility of our results, which can be a true challenge when comparing, for instance, mouse behaviour to human behaviour. Our team at the University of Louisville, headed by my partner, Manuel Casanova, is currently working in that direction and we soon hope to have results which may provide a better framework for the definition of autism.
In terms of genetics, epigenetics, and the like, we also need to take some lessons from the field of cancer research which is at least a decade or more ahead of us. We need to place more efforts into learning from the things other scientific fields have already discovered, because so much of science has been done but is locked behind walls of cross-disciplinary unfamiliarity.
But why reinvent the wheel?
For these reasons, organisations like TWDK are vital in helping to bridge that cross-disciplinary divide. Some of the greatest ideas can be inspired by stepping out of our scientific comfort zones and entertaining new perspectives. That's often when breakthroughs occur.
This guest post is by Emily L. Williams, a Ph.D. candidate in the department of Anatomical Sciences and Neurobiology at the University of Louisville. By day, she studies transgenic mouse models and how various gene products such as Noggin and Pten affect development within the central nervous system, the peripheral nervous system, skin, and cartilage. By night and weekend, she studies autism and analyzes trends in genetics, most recently publishing findings on increased occurrence of transposable elements within high-risk autism genes. On Sundays she also blogs at Science Over a Cuppa, and she can be found @EmLyWill on Twitter. Click here for her full CV.
References |
| How people see autism. Image credit: Hepingting |
Ultimately, that means we study just about everything that appears to have an association with the condition without much understanding of whether the relationship is causal or simply correlational in nature. An excellent example of this comes from studying autism genetics. Over 3,000 genes have found to have minor-to-strong association with the condition, both those which currently have no known cause (idiopathic) and conditions which are characterized by the association of several clinically recognizable features which tend to go together (syndromic), such as Fragile X Syndrome. While all roads may have once led to Rome, not all genes lead to autism. Those thousands of genes represent at least 12% of the genes within the human genome, a staggering proportion. It's highly unlikely that all or most converge onto a single behavioural trait.
just because two things occur frequently together that doesn't mean that one necessarily causes the other. As the hilarious statistical parody teaches us, the loss of pirates does not cause global warming even though the number of pirates have indeed declined over the last 130 years as global temperatures have simultaneously gone up.
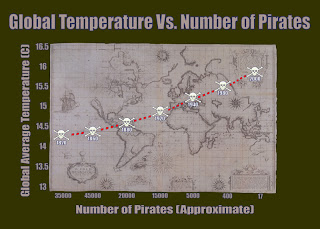 |
| Over the past 130 years, global temperatures have risen, and the number of pirates has decreased. But this does not mean that a lack of pirates causes global warming. Image credit: http://bama.ua.edu |
This is called getting a "false positive", and in fact a recent article in Nature4 just dealt with this exact problem. The researchers reported that some genes which are less stable are more likely to mutate in cancer and therefore appear "associated" even though they are not causally related to the develop of the cancer itself. The problem is when geneticists assume that all genes mutate at the same rate, which is an incorrect assumption. Some genes are more likely to mutate than others, and rates can even vary by person. So the approach to studying mutations requires a more nuanced approach.
As mentioned, thousands of genes have been associated with autism, with approximately 100-200 genes considered "strongly associated". Interestingly, some of the same genes which the Nature paper determined were probably false positives in cancer studies and had nothing to do with the development of the condition are also high up on the autism list. These genes in particular are known for their high mutation rates. Coincidence?
The general point is that cancer researchers have realized that they may be receiving false positives in their genetics studies and are beginning to adjust their methods of analysis to account for this. Given the overlap in important genes associated with both cancer and autism, especially those genes which normally have high mutation rates, autism researchers may need to apply similar methods to whittle down results to those genes most likely play a role in the development of the condition.
 |
| Mice are more similar to us than you might expect. Image credit: Meredith lab, VU University Amsterdam |
In terms of genetics, epigenetics, and the like, we also need to take some lessons from the field of cancer research which is at least a decade or more ahead of us. We need to place more efforts into learning from the things other scientific fields have already discovered, because so much of science has been done but is locked behind walls of cross-disciplinary unfamiliarity.
But why reinvent the wheel?
For these reasons, organisations like TWDK are vital in helping to bridge that cross-disciplinary divide. Some of the greatest ideas can be inspired by stepping out of our scientific comfort zones and entertaining new perspectives. That's often when breakthroughs occur.
This guest post is by Emily L. Williams, a Ph.D. candidate in the department of Anatomical Sciences and Neurobiology at the University of Louisville. By day, she studies transgenic mouse models and how various gene products such as Noggin and Pten affect development within the central nervous system, the peripheral nervous system, skin, and cartilage. By night and weekend, she studies autism and analyzes trends in genetics, most recently publishing findings on increased occurrence of transposable elements within high-risk autism genes. On Sundays she also blogs at Science Over a Cuppa, and she can be found @EmLyWill on Twitter. Click here for her full CV.
why don't all references have links?
[1] Prevalence of Autism Spectrum Disorders — Autism and Developmental Disabilities Monitoring Network, 14 Sites, United States, 2008. Centers for Disease Control and Prevention, MMWR 2012;61(3)
[2] Baron-Cohen S, Lombardo MV, Auyeung B, Ashwin E, Chakrabarti B, et al. (2011) Why Are Autism Spectrum Conditions More Prevalent in Males? PLoS Biol 9(6): e1001081. doi:10.1371/journal.pbio.1001081
[3] Williams et al. (2013). Tranposable elements occur more frequently in autism-risk genes: Implications for the role of genomic instability in autism. Translational Neuroscience, 4(2), 172-202.
[4] Lawrence, Michael S et al. (2013) "Mutational heterogeneity in cancer and the search for new cancer-associated genes." Nature. doi:10.1038/nature12213


No comments:
Post a Comment