We are not alone.
No matter how quietly you listen, nor how closely you stare you’ll not hear them, nor see them with the naked eye. They’re too small, too quiet. They are our microbiome, the trillions of microorganisms that make their homes inside and out of our bodies. They – we – are an ecosystem, with different bugs filling different niches, some helpful, some quietly parasitic, and others, well, others you do end up knowing about…
This is the second of two posts on the subject of the microbiome. The first post looked at questions around the need to unpick its importance to our health – what’s a good or bad microbiome? Which microbes are causes or effects of a disease? And how does the microbiome spread – is it like a disease? Questions which, when answered, could lead to new treatments and ways of protecting our health by manipulating this complex ecosystem within ourselves.
Collateral Damage
Whilst we can’t yet manipulate the microbiome with any finesse, we do influence it through our immune system, evolved over time to cope with invading pathogens and keep them in check. Yet when the immune system isn’t enough, we have a powerful way to, if not manipulate, then affect our microbial ecosystem; it’s just a little…inelegant and unpredictable: Antibiotics.
If the immune system was a sniper, antibiotics would be a bomb, they don’t discriminate between targets. Generally, when you take antibiotics you’re killing off friend and foe alike, this is simply due to how they work; they broadly target bacterial cellular processes, like cell wall building, protein synthesis or metabolism. But their use throws up a number of questions in the light of our growing awareness of the microbiome’s importance. How does the microbiome, as a whole, weather such attacks? How does it recover? And why should we even care?
Well, we know that antibiotic use in early life is associated with a greater risk of developing allergic conditions like asthma1 and irritable bowel disorders2 (IBS) in later life, which is a pretty good reason to care on its own. But how does this happen? Is this due to some interaction of antibiotics on our bodies? It seems pretty unlikely; it’s more likely to be a secondary effect of the ‘collateral damage’ the microbiome takes when we treat an infection.
There’s a growing body of evidence suggesting we rely on the microbiome3, to some extent, to tutor our immune systems. We still don’t know much about the underlying process, but by tampering with the microbiome at an early age we could be messing up our immune systems development, leading to allergic conditions like asthma and IBS.
Weathering it out
So how does the microbiome respond to these antibiotic attacks? In short, we’re not sure, but there are hints.
A study4 by researchers from the University of Valencia followed how the microbiome of a single man, undergoing treatment for an infected pacemaker, faired under antibiotics. Before his treatment they identified 41 species of bacteria, rapidly dropping to 13 by day 11. The total number of species on day 40, after antibiotics, was 38, but all from broadly similar families, lacking the diversity of bugs found on day 0. Why the same families of bugs don’t bounce back, we don’t yet know.
Another problem is that your microbiome’s response to antibiotics may be quite different to mine, and at the moment we can’t predict how much. The microbiomes of different people vary5, there’s no standard ‘set’, so while many groups of bugs share similar biochemical functions and niches, how they respond to antibiotics may differ considerably. We know generally that while under antibiotic attack the microbiome shuts down many of its normal functions, including metabolic processes we rely on, like vitamin production, instead diverting their efforts to protecting themselves, creating defences to fend off the antibiotics.
Like Dominos
Maybe, when it comes to preserving microbial diversity we shouldn’t worry too much about the microbiome as a whole, maybe we could narrow our search a little more.
There is a concept in ecology which is useful to illustrate this: keystone species6. Removing a keystone species can create a domino effect where other species that relied on it will disappear, allowing only a few remainder species to dominate, or new species to invade.
The problem is how, out of so many different species in a microbiome, do we identify a keystone species? Can we predict which ones may survive, and which will perish under a given antibiotic?
There’s good reason to believe antibiotics change the microbiome permanently, and consequentially impact our health. But if we can determine the ‘keystones’ we might be able to target our treatments to avoid causing them too much damage, preserving the ecosystem, as best we can, with minimal detrimental effects, or maybe we could reintroduce them, in the hope of setting our microbiome back on its old path.
This is the second post about the bacteria that live alongside and within us, by freelance science writer Gavin Hubbard. Gavin originally trained as a Medical Biochemist at the University of Surrey and spent over 10 years working in biotechnology, immunology and clinical trials. He writes both for industry and for a general audience, with a focus on health, immunology and pathology. He blogs at Sciencehubb.co.uk and can be found on twitter as @GavinHub
References
why don't all these papers have links?
1. WICKENS, PEARCE, CRANE and BEASLEY (1999), Antibiotic use in early childhood and the development of asthma. Clinical & Experimental Allergy, 29: 766–771. doi: 10.1046/j.1365-2222.1999.00536.x
2. Kronman M.P., Zaoutis T.E., Haynes K., Feng R. & Coffin S.E. (2012). Antibiotic Exposure and IBD Development Among Children: A Population-Based Cohort Study, PEDIATRICS, 130 (4) e794-e803. DOI: 10.1542/peds.2011-3886
3. Arnold I.C., Dehzad N., Reuter S., Martin H., Becher B., Taube C. & Müller A. (2011). Helicobacter pylori infection prevents allergic asthma in mouse models through the induction of regulatory T cells, Journal of Clinical Investigation, 121 (8) 3088-3093. DOI: 10.1172/JCI45041DS1
4. Perez-Cobas A.E., Gosalbes M.J., Friedrichs A., Knecht H., Artacho A., Eismann K., Otto W., Rojo D., Bargiela R., von Bergen M. & Neulinger S.C. (2012) Gut microbiota disturbance during antibiotic therapy: a multi-omic approach, Gut, DOI: 10.1136/gutjnl-2012-303184
5. Schloissnig S., Arumugam M., Sunagawa S., Mitreva M., Tap J., Zhu A., Waller A., Mende D.R., Kultima J.R., Martin J. & Kota K. (2012). Genomic variation landscape of the human gut microbiome, Nature, 493 (7430) 45-50. DOI: 10.1038/nature11711
6. Wagner, S. C. (2012) Keystone Species. Nature Education Knowledge 3(10):51
No matter how quietly you listen, nor how closely you stare you’ll not hear them, nor see them with the naked eye. They’re too small, too quiet. They are our microbiome, the trillions of microorganisms that make their homes inside and out of our bodies. They – we – are an ecosystem, with different bugs filling different niches, some helpful, some quietly parasitic, and others, well, others you do end up knowing about…
This is the second of two posts on the subject of the microbiome. The first post looked at questions around the need to unpick its importance to our health – what’s a good or bad microbiome? Which microbes are causes or effects of a disease? And how does the microbiome spread – is it like a disease? Questions which, when answered, could lead to new treatments and ways of protecting our health by manipulating this complex ecosystem within ourselves.
Collateral Damage
Whilst we can’t yet manipulate the microbiome with any finesse, we do influence it through our immune system, evolved over time to cope with invading pathogens and keep them in check. Yet when the immune system isn’t enough, we have a powerful way to, if not manipulate, then affect our microbial ecosystem; it’s just a little…inelegant and unpredictable: Antibiotics.
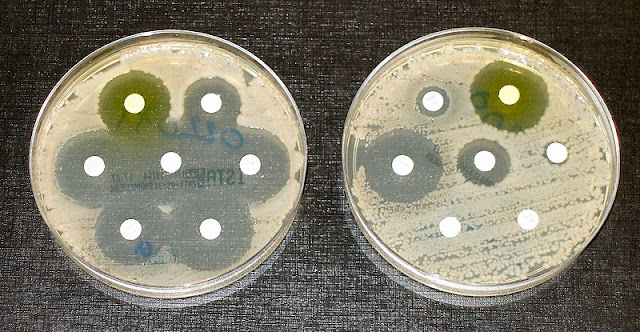 |
| Two different species of bacteria with disks soaked in different antibiotics. The bacteria on the left are susceptible to all the antibiotics tested, while the bacteria on the right are resistant to most of them. Image credit: Wikimedia commons |
If the immune system was a sniper, antibiotics would be a bomb, they don’t discriminate between targets. Generally, when you take antibiotics you’re killing off friend and foe alike, this is simply due to how they work; they broadly target bacterial cellular processes, like cell wall building, protein synthesis or metabolism. But their use throws up a number of questions in the light of our growing awareness of the microbiome’s importance. How does the microbiome, as a whole, weather such attacks? How does it recover? And why should we even care?
Well, we know that antibiotic use in early life is associated with a greater risk of developing allergic conditions like asthma1 and irritable bowel disorders2 (IBS) in later life, which is a pretty good reason to care on its own. But how does this happen? Is this due to some interaction of antibiotics on our bodies? It seems pretty unlikely; it’s more likely to be a secondary effect of the ‘collateral damage’ the microbiome takes when we treat an infection.
There’s a growing body of evidence suggesting we rely on the microbiome3, to some extent, to tutor our immune systems. We still don’t know much about the underlying process, but by tampering with the microbiome at an early age we could be messing up our immune systems development, leading to allergic conditions like asthma and IBS.
Weathering it out
So how does the microbiome respond to these antibiotic attacks? In short, we’re not sure, but there are hints.
 |
| Discoverer of penicillin, Alexander Fleming. Image credit: Public domain |
Another problem is that your microbiome’s response to antibiotics may be quite different to mine, and at the moment we can’t predict how much. The microbiomes of different people vary5, there’s no standard ‘set’, so while many groups of bugs share similar biochemical functions and niches, how they respond to antibiotics may differ considerably. We know generally that while under antibiotic attack the microbiome shuts down many of its normal functions, including metabolic processes we rely on, like vitamin production, instead diverting their efforts to protecting themselves, creating defences to fend off the antibiotics.
Like Dominos
Maybe, when it comes to preserving microbial diversity we shouldn’t worry too much about the microbiome as a whole, maybe we could narrow our search a little more.
There is a concept in ecology which is useful to illustrate this: keystone species6. Removing a keystone species can create a domino effect where other species that relied on it will disappear, allowing only a few remainder species to dominate, or new species to invade.
 |
| In some ecosystems the wolf is a keystone species. If lost, the ecosystem can collapse; their prey – the grazers, such as deer – increase in numbers, eating more foliage, other animals which relied on the foliage either as food or as cover (such as a small rodent) might be more exposed, easier for birds to catch, so they become fewer, and so on. Image credit: Flickr/Creative Commons |
There’s good reason to believe antibiotics change the microbiome permanently, and consequentially impact our health. But if we can determine the ‘keystones’ we might be able to target our treatments to avoid causing them too much damage, preserving the ecosystem, as best we can, with minimal detrimental effects, or maybe we could reintroduce them, in the hope of setting our microbiome back on its old path.
This is the second post about the bacteria that live alongside and within us, by freelance science writer Gavin Hubbard. Gavin originally trained as a Medical Biochemist at the University of Surrey and spent over 10 years working in biotechnology, immunology and clinical trials. He writes both for industry and for a general audience, with a focus on health, immunology and pathology. He blogs at Sciencehubb.co.uk and can be found on twitter as @GavinHub
References
why don't all these papers have links?
1. WICKENS, PEARCE, CRANE and BEASLEY (1999), Antibiotic use in early childhood and the development of asthma. Clinical & Experimental Allergy, 29: 766–771. doi: 10.1046/j.1365-2222.1999.00536.x
2. Kronman M.P., Zaoutis T.E., Haynes K., Feng R. & Coffin S.E. (2012). Antibiotic Exposure and IBD Development Among Children: A Population-Based Cohort Study, PEDIATRICS, 130 (4) e794-e803. DOI: 10.1542/peds.2011-3886
3. Arnold I.C., Dehzad N., Reuter S., Martin H., Becher B., Taube C. & Müller A. (2011). Helicobacter pylori infection prevents allergic asthma in mouse models through the induction of regulatory T cells, Journal of Clinical Investigation, 121 (8) 3088-3093. DOI: 10.1172/JCI45041DS1
4. Perez-Cobas A.E., Gosalbes M.J., Friedrichs A., Knecht H., Artacho A., Eismann K., Otto W., Rojo D., Bargiela R., von Bergen M. & Neulinger S.C. (2012) Gut microbiota disturbance during antibiotic therapy: a multi-omic approach, Gut, DOI: 10.1136/gutjnl-2012-303184
5. Schloissnig S., Arumugam M., Sunagawa S., Mitreva M., Tap J., Zhu A., Waller A., Mende D.R., Kultima J.R., Martin J. & Kota K. (2012). Genomic variation landscape of the human gut microbiome, Nature, 493 (7430) 45-50. DOI: 10.1038/nature11711
6. Wagner, S. C. (2012) Keystone Species. Nature Education Knowledge 3(10):51

Over on Google+ Rajini Rao made a good point about something I had to cut out for reasons of space in the post. Here's the link: https://plus.google.com/u/0/104932769104703486594/posts/9hLGvuwsfcu
ReplyDeleteThe idea was to consider saving a sample of the microbiome from before antibiotic treatment and use it to repopulate the microbiome after treatment is complete. I'm not sure anyone has done this exact thing, but microbiome 'transplants' – effectively a poo transplant –have been used to successfully treat C.diff infections:http://www.nature.com/news/faecal-transplants-succeed-in-clinical-trial-1.12227
https://plus.google.com/u/0/104932769104703486594/posts/9hLGvuwsfcu